By Luciana Hannibal, Ph.D.
Team success goes hand in hand with the professional development of its members. New employees are selected according to background education (know-how theory), previous accomplishments (know-how experience) and potential (know-how to grow). These important metrics help to recruit candidates who are best suited for the position, and capture them at a stage of high self-motivation and drive. Only a few however, will stand out for their achievements, even when the same selection process has been applied to all. Why? A pool of employees will choose to remain in a professional comfort-zone, meeting personal and team demands satisfactorily. Others will strive to further their professional development and to make a difference in the team. Some will go beyond and prepare themselves to lead their own teams in the future. Even after careful recruitment, a population of talented individuals may however derail from successful career development. A mismatch of talents and projects can hamper advancement regardless of individual mindset. Researchers in leading positions are expected to excel at understanding the scientific problem, providing new ideas and solutions, teaching and supervising, cooperating effectively with internal and external colleagues and recruiting copious amounts of extramural funding. Optimizing self-reliance of the team is thus crucial for individual and collective return on investment, and this involves aligning talents with projects. Team leaders must identify individual strengths to effectively assign project roles, and employees must be visible for their most valuable talents. Self-motivated employees, as they typically land in the new job, constitute an invaluable asset.

Here is a brief guide on how to blend projects with selected talent types:
— Published
Categories: Education, Redox Biology, Free Radical Biology and Medicine
Approximately 2.9 million people in the United States suffer from epilepsy, according to the CDC. For patients living with this diagnosis and their doctors it is often difficult to predict the onset or progression of chronic seizures. Thanks to a newly published study from the University of Colorado Skaggs School of Pharmacy and Pharmaceutical Sciences at the Anschutz Medical Campus, that may be changing.
 The study, led by Drs. Manisha Patel and Li-Ping Liang of the University of Colorado, was recently published in Redox Biology, a journal of the Society for Redox Biology and Medicine (SFRBM).
The study, led by Drs. Manisha Patel and Li-Ping Liang of the University of Colorado, was recently published in Redox Biology, a journal of the Society for Redox Biology and Medicine (SFRBM).
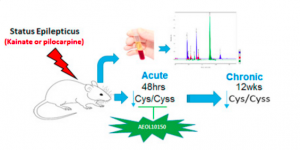
The study was designed to determine if the ratio of reduced and oxidized forms of an amino acid, cysteine and cystine respectively, could serve as an accurate biomarker to predict the onset or progression of seizures associated with epilepsy. Using a rat model, it was determined that decreased cysteine/cystine ratio in plasma may serve as a redox biomarker in epilepsy.
Specifically, seizures were chemically-induced in rats, which were then monitored closely for behavioral changes. Plasma was then taken from the rats 48 hours and 12 weeks after treatment to mimic acute and chronic epileptic conditions. It was found that cysteine/cystine ratio was an accurate redox biomarker for epilepsy. Plasma cysteine/cystine was reduced over 60% in rats with acute epileptic responses and over 37% in rats with chronic epilepsy. Interestingly, cysteine/cystine ratio was unaltered in rats also treated with an antioxidant known to prevent epileptic brain injury.
“Currently the field of epilepsy lacks peripheral blood-based biomarkers that could predict the onset or progression of chronic seizures following an epileptogenic injury,” said Dr. Manisha Patel a Professor at the University of Colorado School of Pharmacy and SFRBM Member. “We are confident that this study is a significant step toward changing this, and will one day help those living with temporal lobe epilepsy.”
About Dr. Patel’s lab
The overarching theme of research in Dr. Patel’s laboratory is to understand the role of redox and metabolic mechanisms in epilepsy. Her laboratory has provided compelling evidence for the role of reactive species and mitochondrial dysfunction in animal models of epilepsy. Furthermore, her research has identified reactive species as novel targets for co-morbidities of epilepsy.
About the University of Colorado Skaggs School of Pharmacy and Pharmaceutical Sciences
Since its inception in 1911, the University of Colorado Skaggs School of Pharmacy and Pharmaceutical Sciences has experienced numerous milestones and is ranked in the top 20 percent of all schools of pharmacy in the country, and fourth in the nation for total NIH funding. Committed to pharmaceutical education, research and patient care, the School educates students in the properties of medicinal agents, the biology of disease, the actions of drugs, and best practices for clinical and therapeutic uses of drugs. Located at the Anschutz Medical Campus, the University of Colorado is the only completely new education, research and patient care facility in the nation today.
— Published
Categories: Free Radical Biology and Medicine, Redox Biology, Research
Click here to read the full study in the journal: Free Radical Biology and Medicine
Cellular redox balance plays a significant role in the regulation of hematopoietic stem-progenitor cell (HSC/MPP) self-renewal and differentiation. Unregulated changes in cellular redox homeostasis are associated with the onset of most hematological disorders. However, accurate measurement of the redox state in stem cells is difficult because of the scarcity of HSC/MPPs. Glutathione (GSH) constitutes the most abundant pool of cellular antioxidants. Thus, GSH metabolism may play a critical role in hematological disease onset and progression.
A major limitation to studying GSH metabolism in HSC/MPPs has been the inability to measure quantitatively GSH concentrations in small numbers of HSC/MPPs. Current methods used to measure GSH levels not only rely on large numbers of cells, but also rely on the chemical/structural modification or enzymatic recycling of GSH and therefore are likely to measure only total glutathione content accurately.
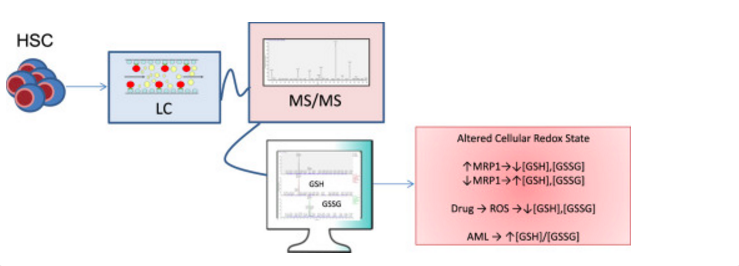
Here, we describe the validation of a sensitive method used for the direct and simultaneous quantitation of both oxidized and reduced GSH vialiquid chromatography followed by tandem mass spectrometry (LC-MS/MS) in HSC/MPPs isolated from bone marrow. The lower limit of quantitation (LLOQ) was determined to be 5.0 ng/mL for GSH and 1.0 ng/mL for GSSG with lower limits of detection at 0.5 ng/mL for both glutathione species. Standard addition analysis utilizing mouse bone marrow shows that this method is both sensitive and accurate with reproducible analyte recovery.
This method combines a simple extraction with a platform for the high-throughput analysis, allows for efficient determination of GSH/GSSG concentrations within the HSC/MPP populations in mouse, chemotherapeutic treatment conditions within cell culture, and human normal/leukemia patient samples. The data implicate the importance of the modulation of GSH/GSSG redox couple in stem cells related diseases.
— Published
Categories: Free Radical Biology and Medicine, Redox Biology, Research

Post by: Peter Vitiello, PhD
Last week, HBO’s satirical comedy newsman, John Oliver, addressed how scientific studies can be misconstrued by the media, transforming their original findings into entertainment solely suitable for morning show banter. In addressing how junk science becomes overhyped, the episode did a great job exposing data replication issues limiting research investments.
(Click Here to View HBO's John Oliver Video Clip. Warning: Language may be inappropriate for some.)
Reproducibility became an important topic of discussion in 2012 when Amgen reported that they were unable to reproduce findings in 47 of 53 “landmark cancer papers” published in journals with an impact factor greater than five. Even when considering failure rates of pre-clinical studies, this 94% irreproducibility rate was shocking and served as the motivation for new standards. The US National Institute of Neurological Disorders and Stroke called for data reporting standards in regard to animal randomization, blinded assessment, sample-size estimation, and data handling and biological sex and reagent authentication have also been added to this list. However, there was also a clear delineation between how this rigor should be applied exploratory studies (early-stage observational tests) and more robust hypothesis-testing experiments.
In light of such issues, direct requests to access raw data and protocol details of published studies have increased. However, editors of the New England Journal of Medicine, which coincidentally has the highest retraction rate of any journal in the world, recently refereed to such scientists as “research parasites”. Casey Greene at the University of Pennsylvania Perelman School of Medicine used this opportunity to create the Research Parasite Award to recognize outstanding contributions to the rigorous secondary analysis of data (nominations are due by October 14). More transparent approaches for handling confirmatory and replicative studies performed by such “parasites” are being developed. Although Amgen’s 47 irreproducible studies remain anonymous, they recently released data on three failed studies through a new “Preclinical Reproducibility and Robustness” channel created by Faculty of 1000. Many other publishers are following suit as Elsevier recently announced a new “Invited Reproducibility Paper” as a new article type appearing in the data science journal, Information Systems.
In summary, I’m very excited to see greater data sharing and transparency through guidelines set by both funding agencies and publishers along with opportunities and respect for parasitic confirmatory studies. I hope that scientists embrace such expectations and do not use them as sole justification to dismiss creativity and novelty during peer review. My only fear is knowing that my harshest critics watch John Oliver and I’ll be expected to respond to these cynics at our next family gathering.
— Published
Category: Redox Biology
On October 21-22, SfRBM leadership held 12 meetings on Capitol Hill in Washington D.C. to introduce key Congressional offices to the society and discuss important issues impacting members, including increased NIH funding, diversity and public engagement in science.
 SfRBM met with senior staff of the Senate Committee on Health, Education, Labor and Pension (HELP), which has jurisdiction encompassing most of the agencies, institutes, and programs of the Department of Health and Human Services including the NIH. SfRBM also met directly with Senator Chuck Grassley, Senator Joni Ernst, Congressman Dave Loebsack to name a few. One goal of these meetings was to build relationships between key Congressional offices and SfRBM. While SfRBM is a member of the Federation of American Society for Experimental Biology (FASEB), which does an amazing job raising awareness of legislation impacting the scientific community, they do not promote or raise awareness about SfRBM specifically.
SfRBM met with senior staff of the Senate Committee on Health, Education, Labor and Pension (HELP), which has jurisdiction encompassing most of the agencies, institutes, and programs of the Department of Health and Human Services including the NIH. SfRBM also met directly with Senator Chuck Grassley, Senator Joni Ernst, Congressman Dave Loebsack to name a few. One goal of these meetings was to build relationships between key Congressional offices and SfRBM. While SfRBM is a member of the Federation of American Society for Experimental Biology (FASEB), which does an amazing job raising awareness of legislation impacting the scientific community, they do not promote or raise awareness about SfRBM specifically.
 SfRBM leaders and staff who participated included Neil Hogg, Rick Domann, Sruti Shiva, Eric Kelley, Kent Lindeman and Brent Carney, the society’s communications and public affairs partner.
SfRBM leaders and staff who participated included Neil Hogg, Rick Domann, Sruti Shiva, Eric Kelley, Kent Lindeman and Brent Carney, the society’s communications and public affairs partner.
In talking to members of Congress about ways to increase NIH research funding, there was pledged support for this initiative, however no real solutions laid out for how to fund it. Congressmen James Sensenbrenner’s office indicated they would not vote in favor of any NIH bill that did not clearly identify the funding source. Senator Robert Casey’s office stated that the pharmaceutical industry, who benefits greatly from the bench science resulting from NIH funding, doesn’t seem to prioritize the importance of NIH funding to support this research when they are speaking with legislators.
 After meeting with the Senate HELP Committee, we came away with guarded optimism. The Committee is currently working on an “Innovation Bill” that will be the Senate companion to the 21st Century Cures Act which passed the House earlier this year. This bill will focus on three sectors, NIH research, FDA research and Health IT. While the bill is set to be introduced in the next couple weeks, making our visit extremely timely, it is unclear if it will contain increased funding for the NIH. Senator Patty Murray, in her role as Committee Ranking Member, indicated to us that such funding is a “must have” in the legislation.
After meeting with the Senate HELP Committee, we came away with guarded optimism. The Committee is currently working on an “Innovation Bill” that will be the Senate companion to the 21st Century Cures Act which passed the House earlier this year. This bill will focus on three sectors, NIH research, FDA research and Health IT. While the bill is set to be introduced in the next couple weeks, making our visit extremely timely, it is unclear if it will contain increased funding for the NIH. Senator Patty Murray, in her role as Committee Ranking Member, indicated to us that such funding is a “must have” in the legislation.
Overall, we were thrilled with the reception we received and the opportunity this gives us in the coming years. Given the timing of the legislation in the HELP Committee and the FY2016 budget, our visit couldn’t have come at a more pivotal time. As one of the benefits you receive as an SfRBM member, we feel this representation in Washington was extremely beneficial for our organization, its members and the issues so many of us hold dear. We look forward to continuing to build the relationships that were started in DC last week, to the benefit SfRBM members in future years.
Sincerely,
Neil Hogg, Ph.D.
Medical College of Wisconsin
SfRBM President
— Published
Category: Redox Biology